Fit3D is a bioinformatics web service to search for spatial residue patterns in proteins, so-called structural motifs. Such motifs play key roles in clarification of linkage between protein structure and function, which is still a demanding process. Fortunately, the comparison of three-dimensional residue patterns can aid the functional comprehension. However, there is an urgent need for up-to-date and versatile computational resources to search for local similarities. Fit3D bridges the gap between easy usability and highly accurate pattern matching. Beside the online version there is also a versatile command line implementation available.
|
The identification and the thorough comprehension of spatial residue patterns can undoubtedly help to bridge the gap between protein structure and function. Spatial arrangements of only a few residues, so-called structural motifs, were observed to play key roles in various biochemical functionalities of proteins like catalytic activity or structure stabilization. Protein-protein or protein-nucleotide interactions are mediated by only a few key residues that can be abstracted to structural motifs as well. Furthermore, it was shown that protein superfamilies can be represented adequately by structural motif templates and hence, distinct homologous structures can be detected. Especially highly diverse protein families like serine proteases are a bottleneck for overall structure comparison methods. | 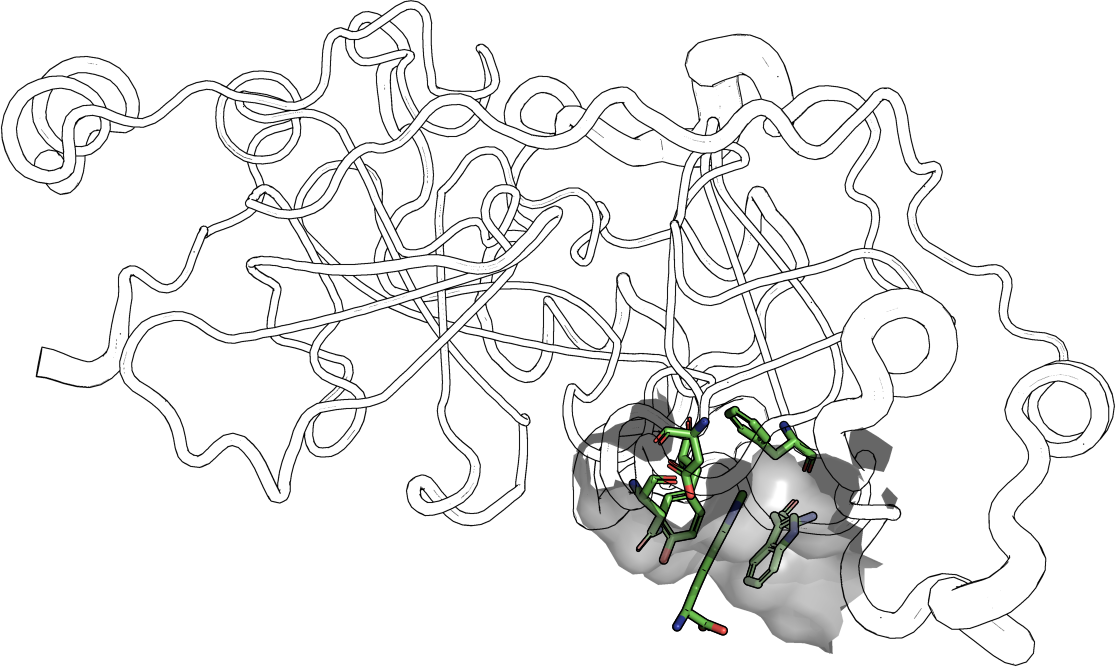 |